Molecule version 2.22 was used throughout the examples and therefore they are not compatible with the recently released version 3 of Molecule.
What is molecule?
No one is perfect. We all make mistakes but we can avoid many of them with careful planning.
Many times I have made changes to an Ansible role and only after going through the process of releasing it to our environment have encountered a simple typo that breaks it, only to fix it, try again and find yet another typo! Or sometimes your role worked fine on Debian but now it doesn’t on CentOS!
Luckily we have a fantastic suite called Molecule to test roles in isolation. Before discovering Molecule I had my own set of test pipelines using Docker and Jenkins but my test scripts were quite rigid and didn’t account for the diversity of configurations and operating systems we manage. It meant my roles were never tested properly.
Using Molecule is much simpler. You can create multiple scenarios for testing configuration variances and you can use Docker images for all the common Operating Systems and versions ensuring they are all thoroughly tested.
Installation
Molecule is written in python and distributed as a pip. In most cases it can be easily installed with the pip command. I am using Docker to test my roles and I therefore need to install the Molecule package with Docker support.
pip install molecule
pip install 'molecule[docker]'If you have trouble with the installation process check the very well documented steps here.
You will also need to have Docker installed and running on your computer.
If you prefer you can run Molecule directly from a Docker image instead of installing it to your system. Quay.io distributes a Molecule Docker image for you to use:
~$ docker run --rm quay.io/ansible/molecule molecule --version
molecule, version 2.20.0.0a3.dev12Init a role
I’m going to make this section quite practical so you can follow through the steps and set up your own roles with Molecule.
First thing is to initialise the role. There are two different ways depending on whether you’re updating an existing role or creating a brand new role.
New Role
The molecule init command creates not just the Molecule directory hierarchy but also the Ansible one.
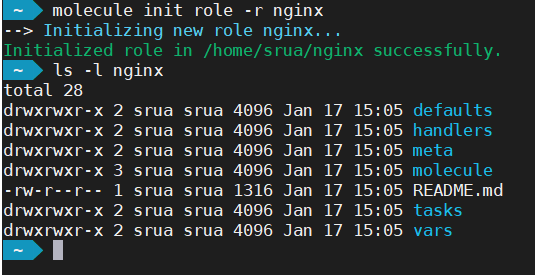
Update Existing
If on the other hand you are updating an existing role, the following command create only the directory molecule with the most basic configuration.
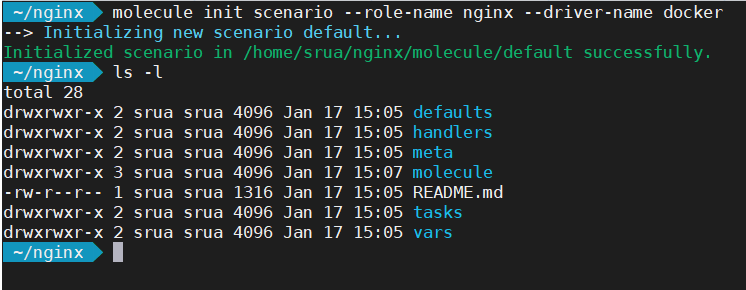
Molecule contents
Let’s first of all examine the contents of the molecule directory:
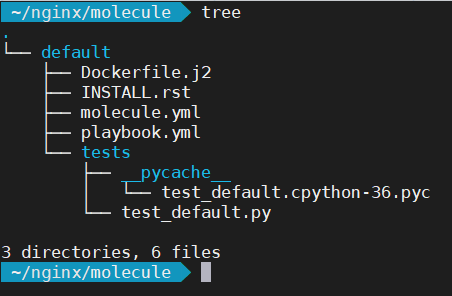
The first thing you’ll notice is there is a default directory. This defines your test scenario. For instance, it allows you to test the same role using different configurations. Inside the default directory there are several files. We’re interested in just the molecule.yml and playbook.yml
Let’s look first at molecule.yml
---
# use Ansible Galaxy to get other modules
dependency:
name: galaxy
# we'll be running our tests inside a docker container
driver:
name: docker
# linter to check file systax; try "molecule lint" to test it
lint:
name: yamllint
# docker images to use, you can have as many as you'd like
platforms:
- name: instance
image: centos:7
# run ansible in the docker containers
provisioner:
name: ansible
lint:
name: ansible-lint
# Molecule handles role testing by invoking configurable verifiers.
verifier:
name: testinfra
lint:
name: flake8And this is playbook.yml which is a standard Ansible playbook to invoke your role.
---
- name: Converge
hosts: all
roles:
- role: nginxYou can add more roles to playbook.yml if you need, variables etc, using standard playbook configs.
First test
You can test your initial setup simply by running molecule create. This command creates the docker images where Ansible will be running your playbooks. Another handy command is molecule login to open a shell into the running docker container for you to perform debugging for example.
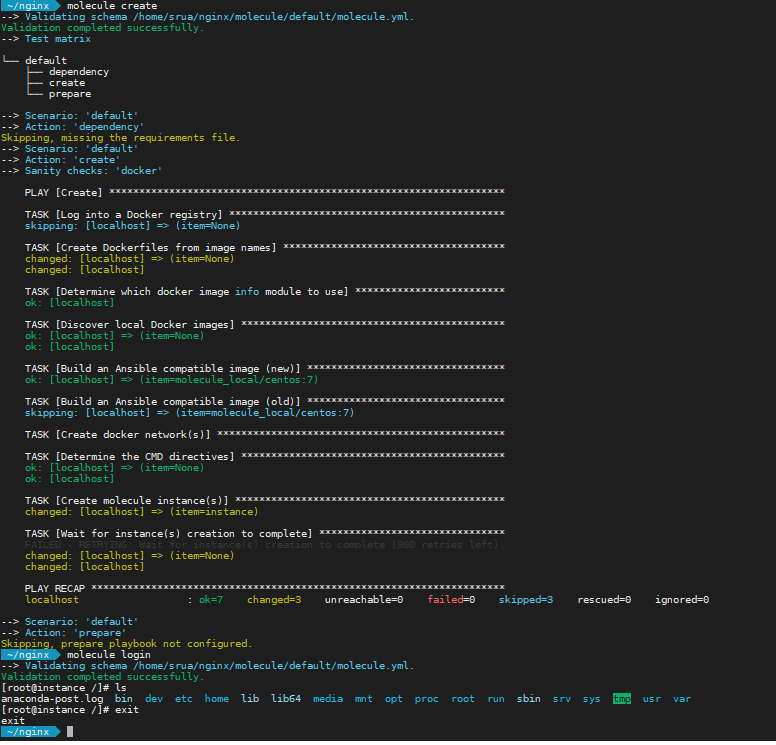
First role
I’m going to assume you are creating a new role for this lab. I’m doing a simple nginx installation. My first change is to make the role install nginx when it runs, adding the following to tasks/main.yml
You may have noticed the default image on molecule is centos7. Feel free to change it to whatever image you prefer but I’m sticking to defaults for the moment and I’ll be showing how to test for multiple distributions shortly.
- name: Install EPEL
package:
name: epel-release
state: present
when: ansible_os_family == "RedHat"
- name: Install nginx
package:
name: nginx
state: presentNow you can simply run molecule converge and the default scenario will run to test your role.
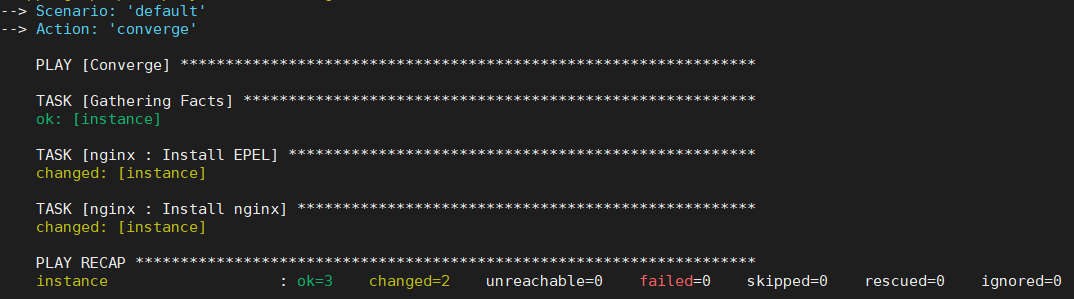
You can use the command molecule test instead which would run through every single step in molecule such as linting, converging, clean up, destroy, etc. But for our tests converge alone is much faster and lint is unlikely to succeed as we haven’t edited the meta/* configurations.
Testing for multiple Operating System
The above role has only been tested for CentOS version 7. But this role may be used on Ubuntu or Debian and we should therefore test them as well. We do this by adding the different images to the platforms section in the molecule.yml file config
platforms:
- name: CentOS-7
image: centos:7
- name: Ubuntu-18
image: ubuntu:18.04molecule converge command will now create two docker images, one for CentOS 7 and one for Ubuntu 18.08 and run Ansible with our brand new role in both.You will need to execute molecule destroy after you change the molecule.yml for Molecule to pick up the new configurations.
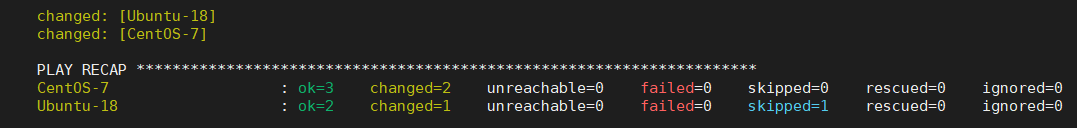
The problem with SystemD
Docker is a containerised system and it can be limiting. It doesn’t always support all the capabilities a host does. I often have problems with some roles requiring config changes to kernel related activities such as sysctl, iptables, etc. In this cases I tend to just ignore the errors by either using the Ansible’s ignore_errors or most often, by setting up a molecule variable and checking for its existence.
- sysctl:
name: vm.swappiness
value: '5'
state: present
when: molecule is not defined
#
# Or
#
- sysctl:
name: vm.swappiness
value: '5'
state: present
ignore_errors: yesBut one of them that is easily fixed is systemd. Most modern GNU/Linux distribution now use systemd for starting and stopping daemons. You will see an error similar to:
TASK [nginx : Start up nginx]
**************************************************
fatal: [CentOS-7]: FAILED! => {"changed": false,
"msg": "Could not find the requested service nginx: "}
changed: [Ubuntu-18]/sbin/init. You do these changes in the molecule.yml as in the example below:platforms:
- name: CentOS-7
image: centos:7
command: /sbin/init
tmpfs:
- /run
- /tmp
volumes:
- /sys/fs/cgroup:/sys/fs/cgroup:romolecule converge it should work. Remember to do a molecule destroy if you’re making configuration changes to Molecule.You can also allow running docker inside Docker by sharing the docker socket with the Molecule containers adding this line to the volumes section:
/var/run/docker.sock:/var/run/docker.sock
Passwords encryption
We use ansible-vault to encrypt all our passwords into the roles we create. There is an argument as to whether roles should have any private information at all (which I agree with) but unfortunately it is sometimes unavoidable for the molecule tests to complete.
Avoid passwords on roles as much as possible. You may want to consider Hashicorp vault as a safer alternative.
This can cause a problem because Molecule will fail if it cannot access the vault. The solution is pretty simple. You just need to edit the molecule.yml and add the path to the vault password file as an argument to Ansible.
provisioner:
name: ansible
log: true
options:
'vault-id': '@$HOME/.ansible/very_secret_file.txt'
lint:
name: ansible-lintRunning from Docker
Just a note to those that decided not install Molecule onto their computer and use Docker instead, your command line is slightly longer but it should work as well:
docker run -ti
-v /var/run/docker.sock:/var/run/docker.sock
-v $(pwd):/nginx quay.io/ansible/molecule
-v /sys/fs/cgroup:/sys/fs/cgroup:ro
/bin/sh -c 'cd /nginx && molecule converge'Continuous Integration
The final piece of the puzzle is the continuous integration. In an ideal world you would not be running Molecule from your own desktop. It makes more sense to automate it using a CI. We prefer Jenkins here as we have an internal git repository but you can also use public CI such as GitLab. You can find examples in the molecule documentation. See below as well:
---
image: docker:git
# uses Docker In Docker
services:
- docker:dind
before_script:
- apk update && apk add --no-cache docker
python3-dev py3-pip docker gcc git curl build-base
autoconf automake py3-cryptography linux-headers
musl-dev libffi-dev openssl-dev openssh
- docker info
- python3 --version
molecule:
stage: default
script:
- pip3 install ansible molecule docker
# you may use "molecule test" to do a full run
# or individual steps if you prefer
- molecule cleanup
- molecule converge
- molecule destroyUsing BitBucket is also pretty straightforward:
---
image: quay.io/ansible/molecule
pipelines:
default:
- step:
deployment: test
services:
- docker
script:
- molecule converge
- molecule destroyA Jenkins example you can use would look like this:
pipeline {
agent {
docker {
image 'quay.io/ansible/molecule'
args '-v /var/run/docker.sock:/var/run/docker.sock -v /sys/fs/cgroup:/sys/fs/cgroup:ro'
}
}
stages {
stage ('Molecule: Clean up environment') {
steps {
sh 'molecule cleanup; sudo molecule destroy'
}
}
stage ('Molecule: run full test') {
steps {
sh 'molecule test --all'
}
}
} // close stages
} // close pipeline- Conclusion
Molecule can be of great help to ensure your roles are up to the best of standards before you tag them for use. It will help you ensuring quality code is used and it works (at least in isolation). It doesn’t mean it will be all perfect, you will rarely be running a single role on your playbook but it should simplify your debugging when problems occur.
But using Molecule alone is not the solution. You should adhere to good DevOps practices and avoid manual process as much as possible. This is what we aim for:
- Jira ticket is created requesting a new feature
- Git branch on the pertinent Ansible roles matching the Jira ticket number is created
- Changes added and committed to the roles
- Pull request initiated
- Our automation (Jenkins) picks up the pull request and runs Molecule on all affected roles
- If Molecule succeeds the pull request can be merge after it has been peered reviewed
- Roles are tag with a version number
Molecule is an integral part of our release process but it would become meaningless if we don’t follow the process.

Sergio Rua
Senior DevOps Engineer
Sergio has many years experience working on various development projects before joining Digitalis. He worked for large companies with complex networks and infrastructure. ‘This has helped Sergio gain lots of experience in multiple areas from programming to networks. He especially excels in DevOps: automation is his day-to-day and Kubernetes his passion.
Related Articles
Getting started with Kafka Cassandra Connector
If you want to understand how to easily ingest data from Kafka topics into Cassandra than this blog can show you how with the DataStax Kafka Connector.
K3s – lightweight kubernetes made ready for production – Part 3
Do you want to know securely deploy k3s kubernetes for production? Have a read of this blog and accompanying Ansible project for you to run.
K3s – lightweight kubernetes made ready for production – Part 2
Do you want to know securely deploy k3s kubernetes for production? Have a read of this blog and accompanying Ansible project for you to run.


